🔴Developed by CDS Empore™
Empore™ E3Technology™
Single-Vessel Proteomics Sample Prep
Protein cleanup, detergent removal, and digestion — all in one column in minutes. Accessible to users of any expertise level, with a cost-effective workflow that eliminates multiple transfer steps.
⚡15-Minute Workflow
🛡 Single Vessel
✓ Detergent Removal


Revolutionary Proteomic Sample Preparation Technology
E3technology™ is an efficient, effective, and economical approach for proteomics sample preparation. Its accessibility makes it suitable for users of any expertise level.
It swiftly conducts protein cleanup, preparing them for digestion in a matter of minutes, while eliminating detergents and interferences. Significantly, it provides a cost-effective solution, and following the cleanup, it seamlessly transitions to the digestion process within the same column.
E3Technology™ Workflow
Save time and improve reproducibility with standardized, pre-packed tips
1
Protein precipitation
Depending on sample volume, add 4x volume of 80% acetonitrile to induce protein precipitation. Note: organic solvents such as cold acetone or 90% methanol can also be used to induce protein precipitation.
2
Sample loading
Transfer protein precipitate to E3products (tip, spin column, cartridge, and plate), centrifuge at 2,000-4,000 rpm for 1-2 min; discard flow through.
3
Wash
Add 200 µl organic solvents, and centrifuge at 2,000- 4,000 rpm for 1-2 min. Discard flow through and repeat this step 2-3 times.
4
Reduction and alkylation
Add 100-500 µl 50 mM triethylammonium bicarbonate (TEAB), 10 mM Tris(2-carboxyethyl) phosphine (TCEP), and 40mM chloroacetamide (CAA), incubate at 45°C for 5 min with gentle shaking.
5
Wash
Centrifuge E3filters at 2,000-4,000 rpm for 1-2 min. Discard flow through. Add 100-500 µl 50 mM TEAB), and centrifuge at 2,000-4,000 rpm for 1-2 min. Discard flow through. The wash step may be repeated 2-3 times in total.
6
Digestion
Transfer E3filters to clean collection tubes, and add 50-200 µl 50 mM TEAB, desired enzyme (Trypsin or Trypsin/Lys-C mix) at a 1:50 ratio. Incubate E3filters at 37°C for 16-18 hours with gentle shaking.
7
Elution
Centrifuge E3filters at 2,000-4,000 rpm for 1-2 min, transfer elutes to new collection tubes. Perform two additional elution steps with 50mM TEAB, and 50% acetonitrile/0.1% formic acid in water, respectively. Pool the elution, dry, and proceed to desalting, or store at -80°C until further use.
Key Features
Revolutionary features that transform proteomics sample preparation
Effective
Compatible with a variety of upstream cell lysis conditions (e.g., urea, SDS, RIPA, TFA, etc.)
Economical
To bridge gaps between genomics and proteomics products
Efficient
<15-min hands-on time
Robust
Zero technical barrier to even entry-level biomedical scientists
Versatile
E3tips™, E3filter™, and E3plate™ to satisfy different sample volumes, concentrations, quantities, and the need for automation.
Stress-free
No liquid transfer, no concerns of any free-beads related issues (protein-bead ratios, beads sticking to tube walls and surfaces, cross-contaminations, etc.)
E3Technology™ Performance
Quantitative results demonstrating superior protein identification and reproducibility
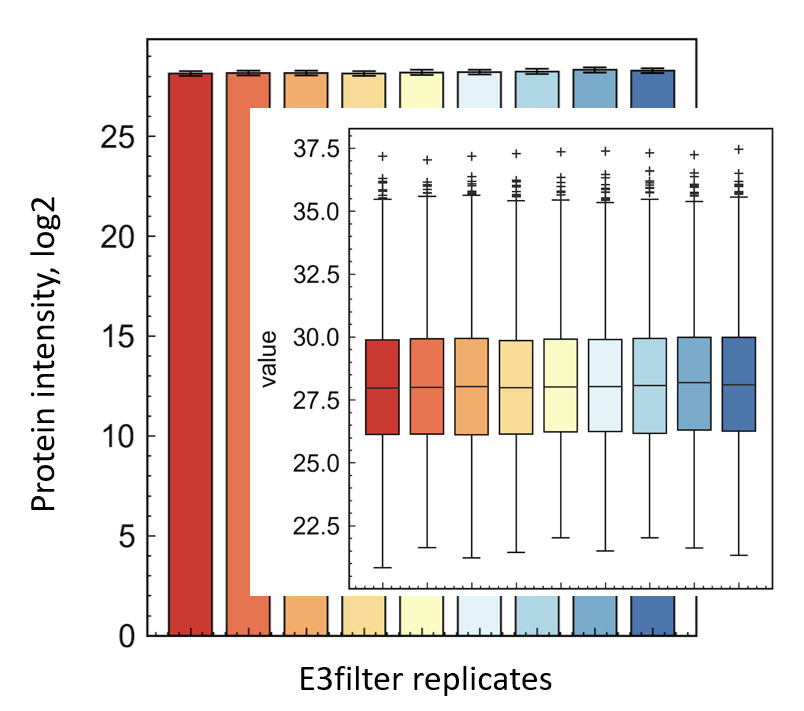

.jpg)



Application Notes

Figure 1. Qualitative assessment of E3technology (E3filter) for E. coli proteome analysis
(A-C) Comparison of the number of proteins, peptides, and PSMs between the E3filter, FASP, and SP4-GB approaches. Error bars represent three replicates.
(D-E) Overlapping analyses of proteins and peptides derived from the three methods

Figure 2. Qualitative assessment of E3technology (E3filter) for E. coli proteome analysis
(A-C) Comparison of the number of proteins, peptides, and PSMs between the E3filter, FASP, and SP4-GB approaches. Error bars represent three replicates.
(D-E) Overlapping analyses of proteins and peptides derived from the three methods
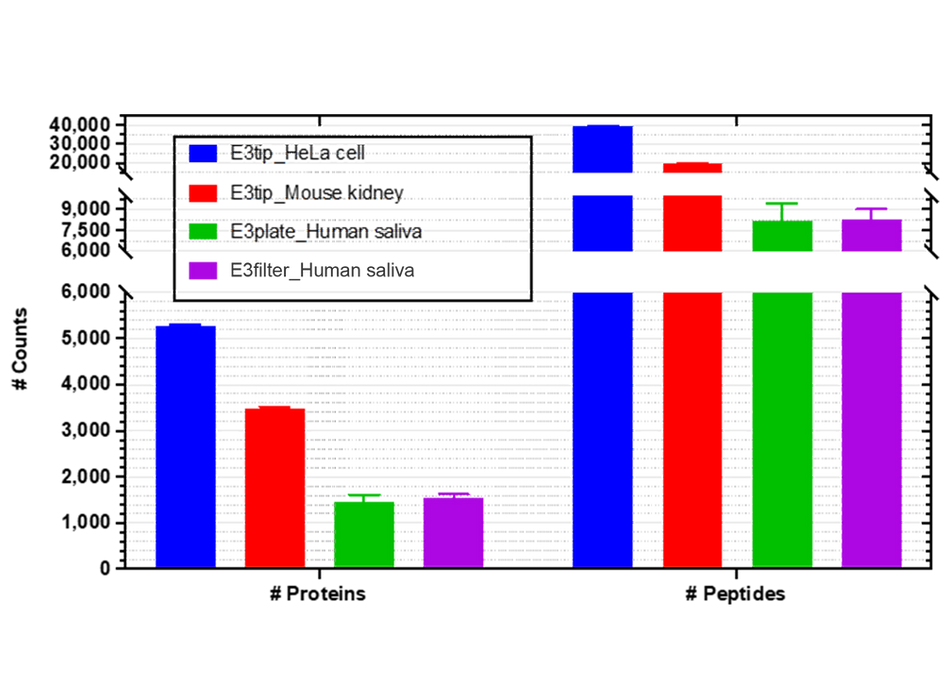
Figure 3. Qualitative assessment of E3technology (E3filter) for E. coli proteome analysis
(A-C) Comparison of the number of proteins, peptides, and PSMs between the E3filter, FASP, and SP4-GB approaches. Error bars represent three replicates.
(D-E) Overlapping analyses of proteins and peptides derived from the three methods
Products Selection Guide
Choose the right E3 procducts for your application

